Researchers take cues from nature to form living materials with unprecedented control and versatility.
October 22, 2019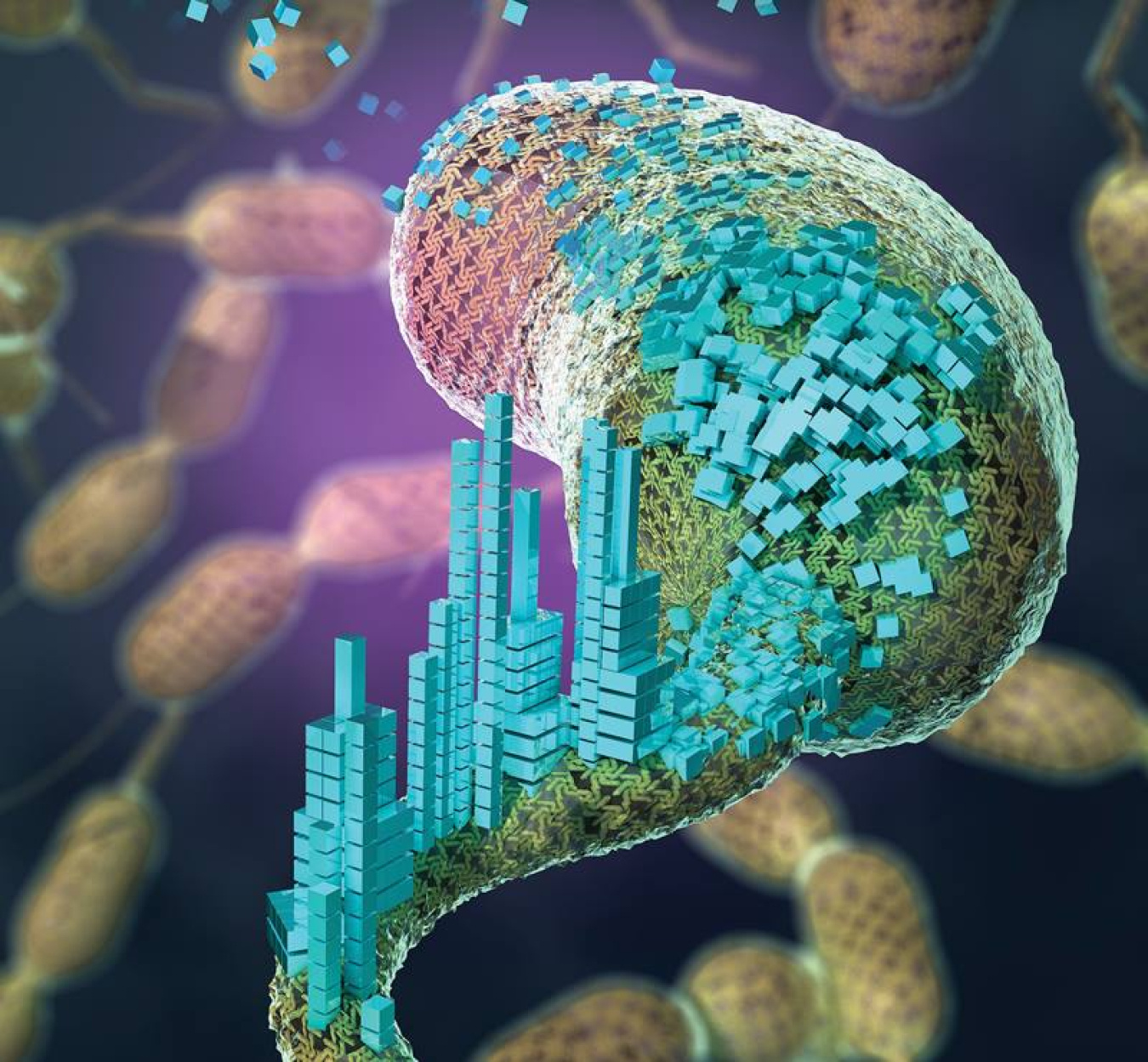
The Science
Bone and mollusk shells are composite systems that combine living cells and inorganic components. This allows them to regenerate and change structure while also being very strong and durable. Borrowing from this amazing complexity, researchers have been exploring a new class of materials called engineered living materials (ELMs). These materials combine living cells and synthetic components. In addition, they use the living cells as “materials scaffolds” that can organize or modify the material throughout its lifetime. In particular, researchers have engineered a set of bacteria that can irreversibly attach hard or soft materials to a cell surface.
The Impact
Researchers have created a living construction kit. This new approach can make materials with distinct structures at each size level. It also has the highest surface density of any other engineered living material (ELM). Unlike previous approaches, scientists can control how many proteins are attached to the surface and where they are placed. The new platform opens the door to creating new applications in bioelectronics, biosensing, smart materials, and catalysis.
Summary
Both natural structures and the ELMs they inspire are made up of hierarchical patterns of materials across multiple length scales. This means that for a material made of regularly sized building blocks, each block is made of smaller blocks, and each of the smaller blocks is made of even smaller pieces. For example, mollusks build their shells out of layers of calcium carbonate that are millimeters (mm) thick. Each layer is made of calcium carbonate platelets that are 500 nanometers (nm) thick and about 5–10 microns (micrometers or µm) wide. Finally, each platelet is made of millions of nanograins (each only 30 nm in size) glued together by a biopolymer. Thus, within a single shell, there are distinct structures at several different length scales (10’s of nm, 100’s of nm, µm, and mm).
Earlier approaches to designing ELMs relied on 1-D protein filaments (like strings) or isolated membrane proteins on cell surfaces to attach materials to a bacterium. The limitation to this strategy is that there is little control over where these proteins are placed on the surface or how tightly packed together they are. In this new work, the researchers targeted surface layer (S-Layer) proteins since they form ordered, sheet-like structures on the surface of many microbes.
Specifically, the researchers chose to use RsaA, a well-studied S-Layer protein with a known atomic structure. It is from the bacterium Caulobacter crescentus, a microorganism that can survive under low-nutrient and low-oxygen conditions, which is ideal for the long-term survival of an ELM.
In order to use the S-Layer of C. crescentus to control material assembly, the researchers employed a pair of proteins that work like a biological lock and key in a system called SpyTag/SpyCatcher. SpyTag peptides (chains of amino acids) are inserted into the RsaA S-Layer of a cell and will only bond to materials that carry its SpyCatcher partner peptide. This approach offers a versatile yet precise platform for ELMs, as it allows any material with SpyCatcher to be bonded to the cell surface at specific positions.
The researchers discovered that they could control how much material could be attached to the cell by changing the location of the SpyTag peptides. The density of attached material can be tuned from 4% of the RsaA lattice all the way to 23%, which is the highest density of cell surface coverage reported for existing ELMs. Using this system, the researchers found that the engineered bacteria could attach proteins, biopolymers, and quantum dots to the engineered cell surface and that these modifications did not affect the cells' viability.
Contact
Caroline Ajo-Franklin
Molecular Foundry, Berkeley Lab
Currently at Rice University
caroline.ajo-franklin@rice.edu
Funding
This work was supported by the Defense Advanced Research Projects Agency (Engineered Living Materials Program, C.M.A-F.) and National Institutes of Health award R01NS096317 (B.E.C.) Work at the Molecular Foundry was supported by the Office of Science, Office of Basic Energy Sciences, of the U.S. Department of Energy under Contract No. DE-AC02-05CH11231.
Publications
M. Charrier, et al. “Engineering the S-Layer of Caulobacter crescentus as a Foundation for Stable, High-Density, 2D Living Materials." ACS Syn. Bio. (2019). 8. 181-190. DOI: 10.1021/acssynbio.8b00448

