Soils contain a large diversity of unknown RNA viruses that infect fungi and possibly plants and animals.
Biological and Environmental Research
May 7, 2020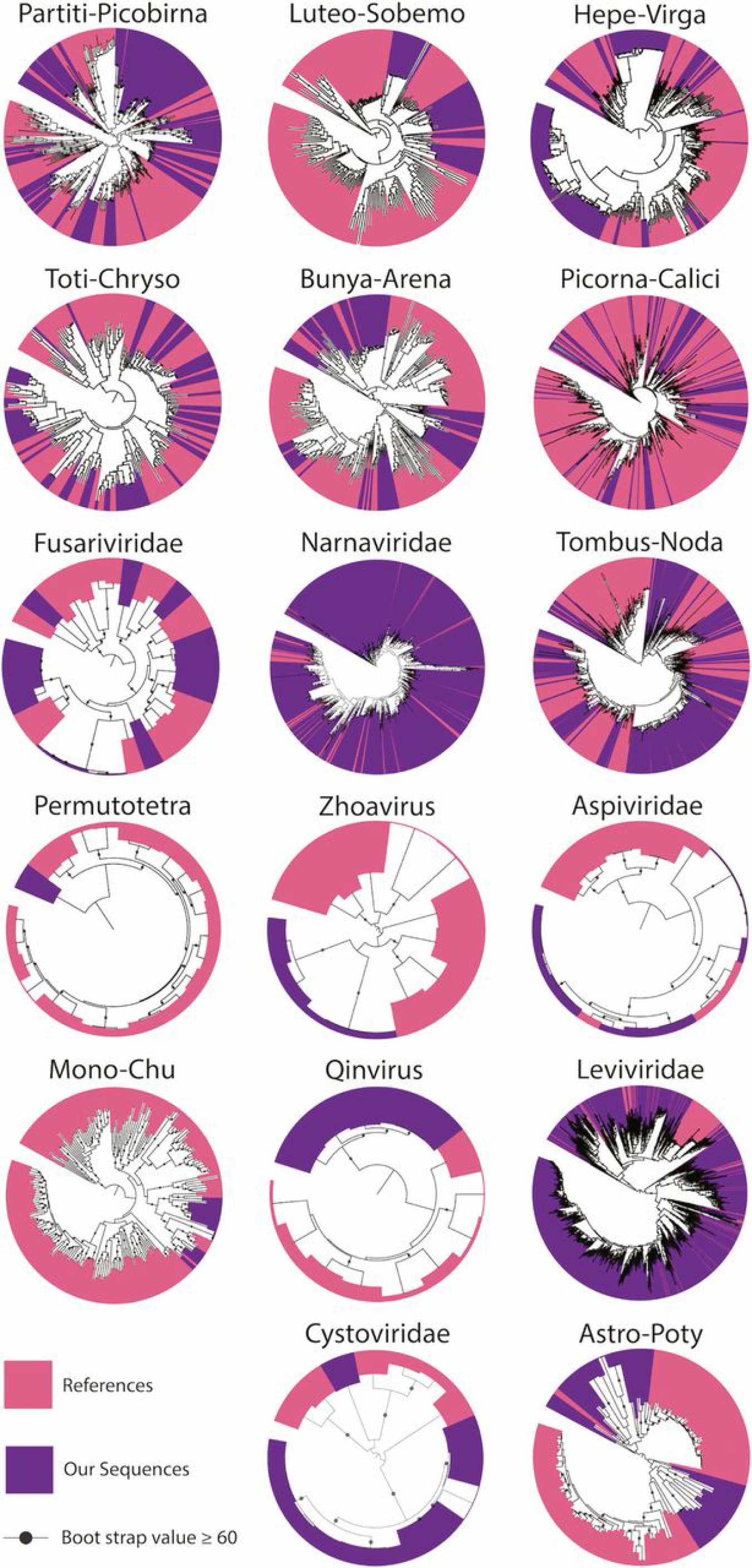
The Science
Viruses play a significant role in the ecology of the oceans. For example, they shape the diversity of natural bacteria populations. Viral activity can change the distribution of elements in the environment and cause blooms of microbes to collapse. In contrast to the oceans, little is known about the role of viruses in soils. A new study found that soils can contain many kinds of RNA viruses. Most of these RNA viruses likely infect fungi, but they could also infect bacteria, plants, and animals. The study found that viral populations in soil change quickly. This means viruses may be multiplying and responding to environmental changes.
The Impact
The study is the first example of a sequencing-based analysis of RNA viruses in soil. Sequence analysis involves the use of multiple, computational methods to understand a genetic sequence’s features, function, structure, or evolution. The results suggest that soil RNA viruses are diverse, abundant, and infect primarily fungi and some bacteria. However, they possibly also infect vertebrates and plants. These data suggest viruses are potentially as important in the turnover of chemicals in soils as they are in the ocean. Understanding the genetic makeup and diversity of viruses in soils may help us control the microorganisms in soils and may reveal new genetic features with applications in biotechnology.
Summary
Viruses are the largest genetic reservoir on Earth. They have important implications for agriculture and public health, and scientists believe viruses to be major drivers of biogeochemical processes in the soil environment. Biogeochemical processes involve the turnover from one chemical form to another. Virtually nothing is known about RNA viruses in soils, despite their widespread distribution and the profound importance of soils to human activities. In this study, scientists sampled controlled soil samples over a time series and analyzed what genes the microbes actively used. The aim was to understand the diversity of soil viruses and how their communities change over time. The analysis indicates that the most abundant viruses were part of the family Narnaviridae and likely infect fungi. The study also identified a second major group of viruses, the Leviviridae, which may infect Proteobacteria. Viral and host community structure was dynamic and responded to experimental treatments with root litter. This suggests that viral communities were actively replicating and responding to environmental change. These findings expand our understanding of the importance of RNA viruses in the environment and have important implications for carbon cycling in soils.
Contact
Principal Investigator
Dr. Mary K. Firestone
Department of Plant and Microbial Biology
University of California, Berkeley
mkfstone@berkeley.edu
Program Manager
Dr. Dawn Adin
Department of Energy (DOE)
Office of Science (SC)
Office of Biological and Environmental Research (BER)
Biological Systems Science Division (BSSD)
dawn.adin@science.doe.gov
Funding
This work was supported by the Department of Energy Office of Science, Office of Biological and Environmental Research via grants to M.K. Firestone (SC0010570 and SC0016247), J. Pett-Ridge (SCW1589 and SCW1632), and J.F. Banfield (SC0010566). E. Starr received additional support through a National Science Foundation fellowship. Sequencing was conducted as part of Community Sequencing Awards 1487 (to J. Pett-Ridge) and 1472 (to M.K. Firestone) by the US Department of Energy Joint Genome Institute, a Department of Energy Office of Science User Facility.
Publications
Starr, E., Nuccio, E., Pett-Ridge, J., Banfield, J. & Firestone, M., “Metatranscriptomic reconstruction reveals RNA viruses with the potential to shape carbon cycling in soil.” Proceedings of the National Academy of Sciences 116, 25900-25908 (2019). DOI: 10.1073/pnas.1908291116.

