The subterranean world hosts up to one fifth of all biomass, including microbial communities that drive transformations central to Earth’s
Biological and Environmental Research
January 23, 2017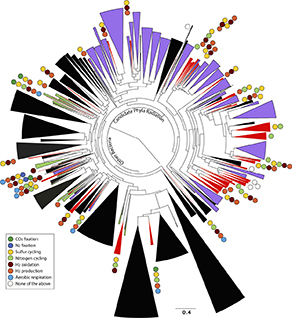
The Science
One of the most detailed genetic studies of any ecosystem to date has uncovered incredible biological diversity among subsurface bacteria. Researchers reconstructed the complete sets of genetic material, or genomes, of more than 2,500 microbes. The team took these microbes from sediment and groundwater samples collected from an area prone to flooding in Colorado. These genomes represent 80 percent of all known bacterial phyla. Analyses showed that interorganism interactions are required to turn the carbon, sulfur, and nitrogen cycles. Further, analyses revealed that complex patterns of community assembly are likely key to ecosystem functioning and resilience.
The Impact
This research has nearly doubled the number of known bacterial groups. It has also provided detailed information about the ecosystem roles of organisms from these groups. Additionally, this research has dramatically increased our understanding of subsurface biology. It will motivate new approaches to ecosystem modeling. The genomes themselves represent a treasure trove that scientists will mine for biotechnology, where bacteria and other organisms are applied to recycle wastes or perform other tasks.
Summary
The subterranean world hosts up to one fifth of all biomass, including microbial communities that drive transformations central to Earth's biogeochemical cycles. However, little is known about how complex microbial communities in such environments are structured and how interorganism interactions shape ecosystem function. A team of researchers, led by Lawrence Berkeley National Laboratory and the University of California, Berkeley, applied terabase-scale cultivation-independent metagenomics to aquifer sediments and groundwater samples and reconstructed 2,540 high-quality, near-complete, and complete strain-resolved genomes that represent the majority of known bacterial phyla. Some of these genomes derive from 47 newly discovered phylum-level lineages. The scientists used metabolic analyses spanning this vast phylogenetic diversity and representing up to 36 percent of the organisms detected in the system to document the distribution of pathways in coexisting organisms. Consistent with prior findings indicating metabolic handoffs in simple consortia, this research shows that a few organisms within the community conduct multiple sequential redox transformations. As environmental conditions change, different assemblages of organisms are selected to alter linkages among the major biogeochemical cycles.
Contact
BER PM Contact
David Lesmes, SC-23.1, 301-903-2977
Contact
Susan Hubbard
Lawrence Berkeley National Laboratory
sshubbard@lbl.gov
Funding
This work was funded by the U.S. Department of Energy (DOE), Office of Science, Office of Biological and Environmental Research, Subsurface Biogeochemical Research program through Lawrence Berkeley National Laboratory's Sustainable Systems Scientific Focus Area. Terabase-scale sequencing was provided by DOE's Joint Genome Institute via Community Science Program allocations.
Publications
K. Anantharaman, C.T. Brown, L.A. Hug, I. Sharon, C.J. Castelle, A.J. Probst, B.C. Thomas, A. Singh, M.J. Wilkins, U. Karaoz, E.L. Brodie, K.H. Williams, S.S. Hubbard, and J.F. Banfield, "Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system." Nature Communications 7, article 13219 (2016). [DOI: 10.1038/ncomms13219]
Highlight Categories
Performer/Facility: University, DOE Laboratory, SC User Facilities, BER User Facilities, JGI

