KBase is an open-source, extensible community resource that enables data sharing, integration, and analysis of genomic information of microbes,
Biological and Environmental Research
August 22, 2018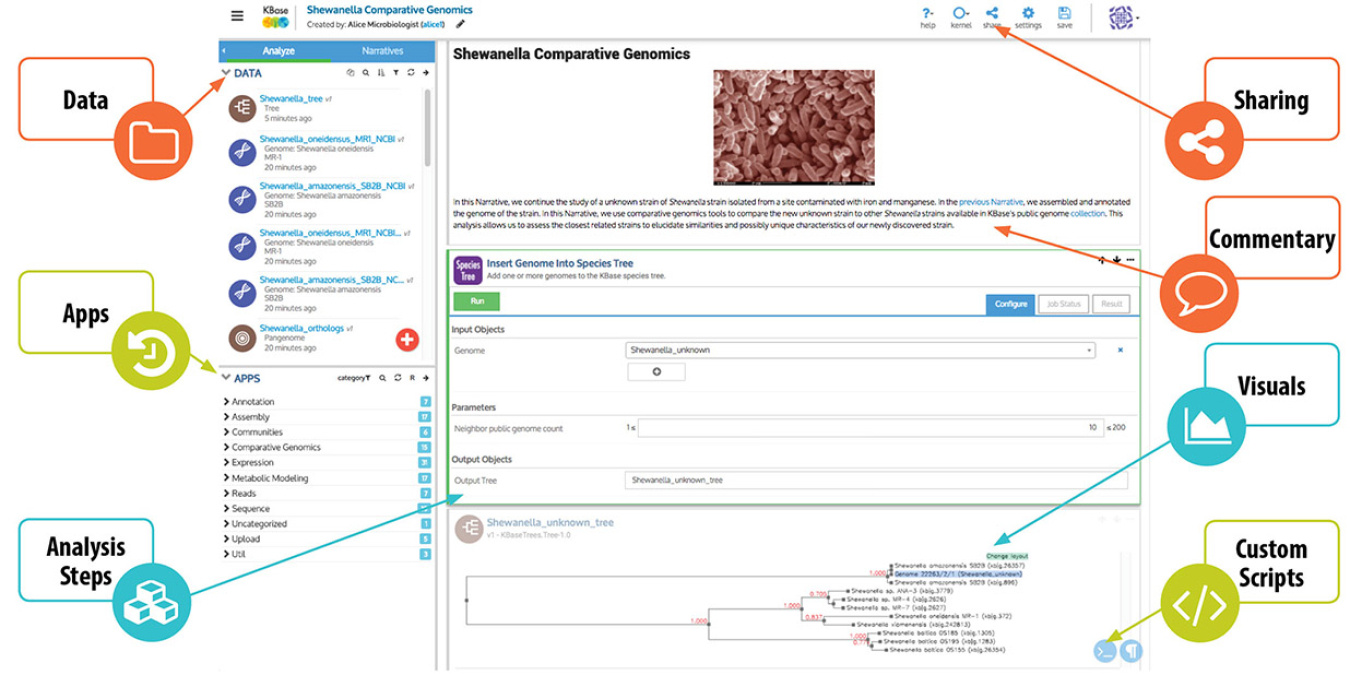
In KBase, anyone can upload and analyze their data and share their results and commentary with other researchers, who can rerun or build upon their work, accelerating the pace of scientific progress.
The Science
Advance biofuel and bioproduct production. This is a key goal of Department of Energy (DOE) research. We need to further our understanding of plants and microbial systems as a basis for developing innovative processes for bioenergy and bioproducts production from inedible cellulosic biomass. To accelerate research, scientists need to perform collaborative, complicated analyses. They must have access to very large and disparate datasets. The DOE Systems Biology Knowledgebase (KBase) is a free, open-source software and data platform that meets these needs. It lets researchers collaboratively generate, test, compare, and share hypotheses about biological functions. It makes research accessible, shareable, and reproducible.
The Impact
KBase empowers scientists across a broad range of systems biology domains that involve big data and require high-performance computing. These domains include environmental analysis, biosystems design, and bioenergy. Science done so far within KBase appears in over 40 peer-reviewed articles. Scientists have used KBase to analyze large microbial communities, predict plant-microbe interactions, and model the metabolism of microbes. The KBase Narratives associated with these experiments are available. They can be copied, re-run, and extended by others.
Summary
KBase is an open-source, extensible community resource that enables data sharing, integration, and analysis of genomic information of microbes, plants, and their communities. KBase's growing suite of scientific tools and reference data offers "one-stop shopping" for users who want to build and share sophisticated bioinformatics workflows. For example, a user can predict species interactions from metagenomic data by assembling raw DNA sequencing reads, binning assembled contigs (sets of overlapping DNA segments that together represent a consensus region of DNA) by species, annotating genomes from these bins, and reconstructing and analyzing individual and community-level metabolic models based on these genomes. External developers can add open-source analysis tools to KBase to make them available to all and allow users to choose among different tools that may have different strengths for particular datasets or workflows. KBase's Narrative Interface, built on the Jupyter platform, lets users do the following:
- Upload their data
- Search and retrieve extensive public reference data
- Access data shared by others
- Share their data with others
- Select and run applications on their data
- View and analyze the results from those applications
- Record their thoughts and interpretations along with the analysis steps.
All of this work is saved in the Narratives, which are private by default but users can choose to make their Narrative public or share it with other individual users. Recording a user's KBase activities in a sharable Narrative is a central pillar of KBase's support for reproducible, transparent research.
Contact
Adam P. Arkin
Principal Investigator
Lawrence Berkeley National Laboratory
aparkin@lbl.gov
Robert Cottingham
Co-Principal Investigator
Oak Ridge National Laboratory
cottingham@ornl.gov
Christopher Henry
Co-Principal Investigator
Argonne National Laboratory
chenry@anl.gov
Funding
This work is supported by the Department of Energy, Office of Science, Office of Biological and Environmental Research Genomic Science Program.
Publications
A.P. Arkin, R.W. Cottingham, C.S. Henry, et al., "KBase: The United States Department of Energy Systems Biology Knowledgebase." Nature Biotechnology 36, 566 (2018). [DOI: 10.1038/nbt.4163]
Related Links
KBase website: http://kbase.us
KBase publications list: http://kbase.us/publications

