A new computational method reliably predicts interactions that depend on neighboring organisms in an environment.
Biological and Environmental Research
December 20, 2019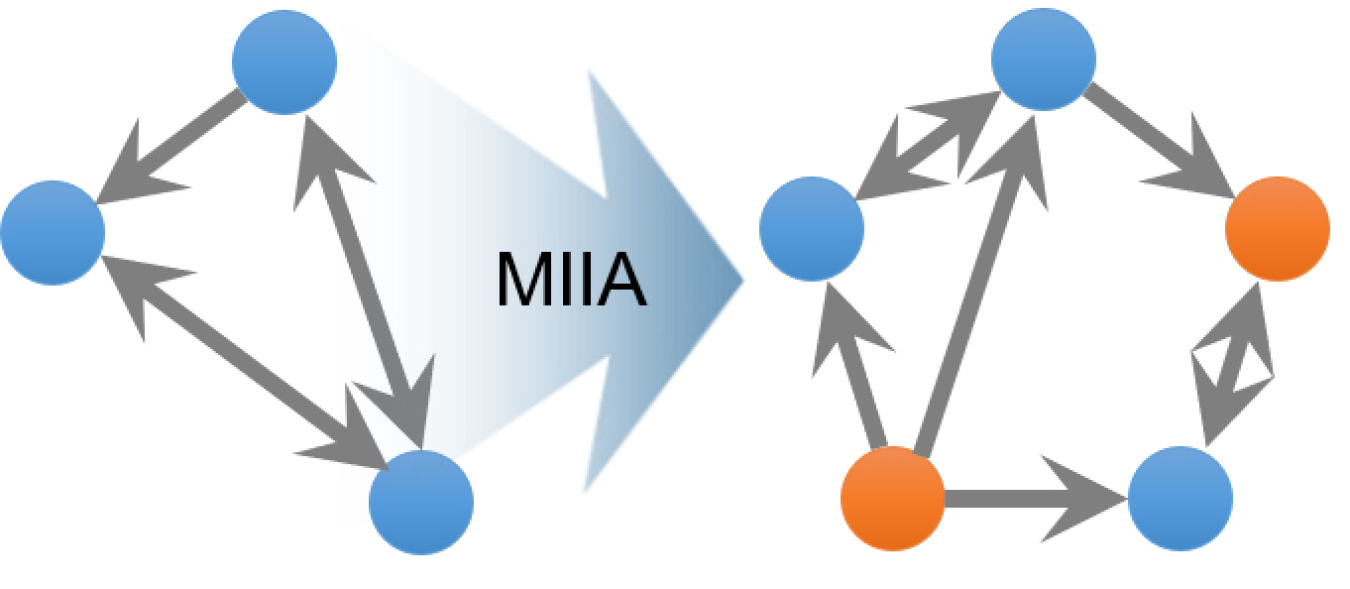
The Science
Microbes in the soil form networks, which in turn make up larger communities. As the environment changes, so do the microbes and their relationships. For scientists to design or control these microbes, they must first understand their communities in nature (microbiomes). Soil ecologists know well that neighboring species influence some microbial interactions. Researchers developed a new theoretical framework called minimal interspecies interaction adjustment (MIIA). It predicts how surrounding organisms and other factors drive changes in interactions in microbial communities.
The Impact
This new computational method improves our understanding of how microbes organize themselves into communities. It describes in detail how neighboring species affect interactions between microbes. This method also predicts major shifts in microbes’ influence on carbon and nitrogen cycles. This information could enable scientists to design and engineer groups and communities of microbes in the future.
Summary
Microbial community dynamics in soil and other habitats involve nonlinear interspecies interactions, so these dynamics are notoriously difficult to predict. Yet understanding how such microbiomes are organized in nature is necessary for designing them (such as for biofuel production) and for controlling them—for example, as a way to assure that soils do not emit too much carbon into the Earth’s atmosphere. Meanwhile, ecologists know that interactions in microbial communities are influenced by neighboring species, or what organisms are around them. Until now, however, there has been no theoretical framework that can predict such context-dependent microbial interactions.
The research was motivated by the following fundamental ecological questions: How are interspecies interactions modulated by shifts in community composition and species populations? To what extent can interspecies relationships observed in simple cultures be translated into complex communities?
The researchers addressed these questions by demonstrating that the theoretical framework enables microbial interactions in binary, or one-to-one cultures to be translatable into complex communities. The researchers also demonstrated the utility of this method in designing and engineering microbial consortia. In this regard, they found that microbial interactions can be significantly modulated when perturbed by a small number of neighboring species—but that the level of modulation diminishes as the number of new neighboring species increases.
This work, the authors say, can also be applied to questions of community ecology beyond microbes. It may provide a theoretical platform for better understanding all biological interaction systems.
Contact
Contact
Hyun-Seob Song
Pacific Northwest National Laboratory
HyunSeob.Song@pnnl.gov
Janet Jansson
Pacific Northwest National Laboratory
janet.jansson@pnnl.gov
Program Manager
Dawn Adin
DOE Office of Biological and Environmental Research, Biological Systems Science Division
Dawn.Adin@science.doe.gov
Funding
This research was supported by the U.S. Department of Energy (DOE) Office of Biological and Environmental Research (BER), as part of Foundational Scientific Focus Area (SFA), Soil Microbiome SFA, and Subsurface Biogeochemistry Research (SBR) SFA at the Pacific Northwest National Laboratory (PNNL).
Publications
H-S. Song, J-Y. Lee, S. Haruta, W.C. Nelson, D-Y. Lee, S.R. Lindemann, J.K. Fredrickson, and H.C. Bernstein, “Minimal Interspecies Interaction Adjustment (MIIA): Inference of Neighbor-Dependent Interactions in Microbial Communities,” Frontiers in Microbiology (2019). [DOI: 10.3389/fmicb.2019.01264]

