Scientists get a molecular look at how plants and bacteria interact, with insights for agriculture.
Biological and Environmental Research
June 9, 2021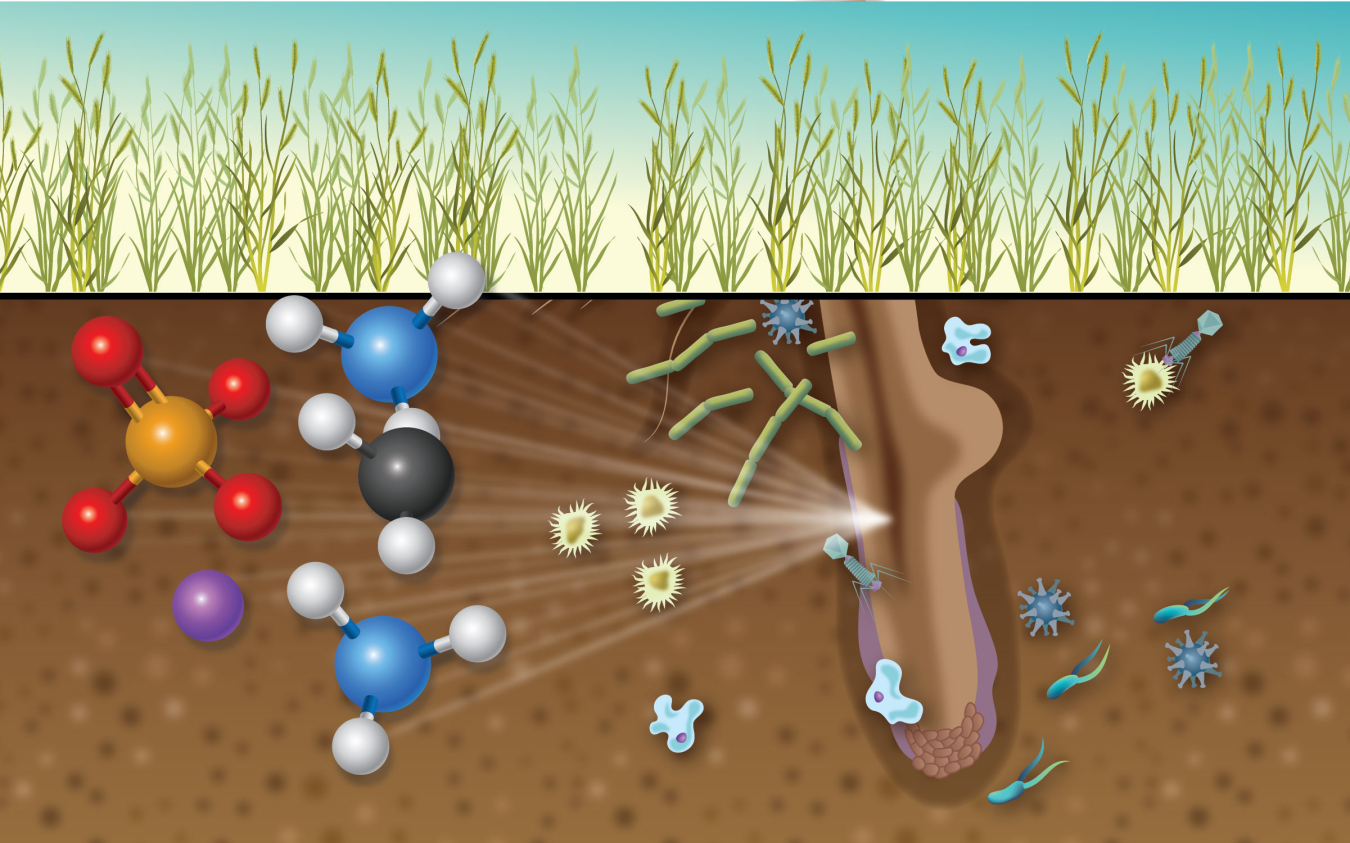
For the first time, scientists viewed the interactions between bacteria and plants at the molecular level, offering insights for improving agriculture.
The Science
The rhizosphere—the thin region of soil surrounding and including the roots of plants—is a well-known hotspot for microbes like bacteria. These bacteria serve a host of beneficial functions for plants. They can reduce infections, enhance tolerance to drought, and help plants take up key nutrients through their roots. However, scientists did not fully understand how the bacteria assist plants. Using four different instruments and a universal sample holder, a new study provides unique insights into the spots on root segments where bacteria attach. These spots are also where special carbohydrate polymers form and where sodium, sulfur, and other organic species are enriched.
The Impact
The rhizosphere may be the most complex microbial habitat on Earth. If scientists could understand, predict, and control interactions between plants and bacteria there, they could harness these interactions to increase or restore plant productivity. Scientists could also use bacteria as biofertilizers or natural pesticides. That could reduce pollution from chemical fertilizers and toxic pesticides. Bacteria could even form part of ecosystems for long-term soil carbon storage to help mitigate the effects of a changing climate. The results of the new study build the foundation to understand these interactions better. They also contribute to the basic science of microbiology.
Summary
Scientists from the Environmental Molecular Sciences Laboratory (EMSL), a Department of Energy Office of Science user facility, joined with colleagues from other organizations to better understand microbial interactions with plant roots within the rhizosphere. Previous approaches to study this below-ground region lacked the high spatial resolution of chemical imaging at the molecular level, which is necessary to capture interactions at the plant-microbe, microbe-microbe, and plant-plant interfaces. The team reasoned that complementary instruments might produce a more useful image at this very small scale if the instruments could be pointed to the same tiny spot across each sample. To accomplish this, they designed an imaging sample holder that allows integrated surface imaging tools to map the same locations of a plant root-microbe interface at the molecular level. The approach—which used EMSL’s time-of-flight secondary ion mass spectrometry, fluorescence microscope, high resolution microprobe XPS, and a scanning electron microscope—was the first to yield a detailed, composite image of plant root-bacterial interactions. This biomolecular imaging data suggest that the root surface is more uneven than expected, with the interaction between the root and bacteria confined to only a few spots along the sampled root segments. The researchers also found these spots to be enriched in sodium, sulfur, and high-mass organic species. The results could help guide future research into the inner workings of the rhizosphere. This work was performed as part of EMSL’s Molecular Plant Phenotyping Integrated Research Platform.
Contact
BER PM Contact
Paul Bayer, Department of Energy Office of Science
Paul.Bayer@science.doe.gov
PI Contact
Zihua Zhu
Environmental Molecular Sciences Laboratory
zihua.zhu@pnnl.gov
Funding
This work was supported by the Department of Energy Office of Science, Office of Biological and Environmental Research, including support of EMSL, a DOE Office of Science user facility.
Publications
Liu W., et al., Correlative surface imaging reveals chemical signatures for bacterial hotspots on plant roots. Analyst 145, 393–401 (2020) (Inner Front Cover). [DOI: 10.1039/c9an01954e]

