For more than half of the world’s population, rice is the primary staple crop. As a grass, it is a close relative of the candidate bioenergy
Biological and Environmental Research
October 8, 2017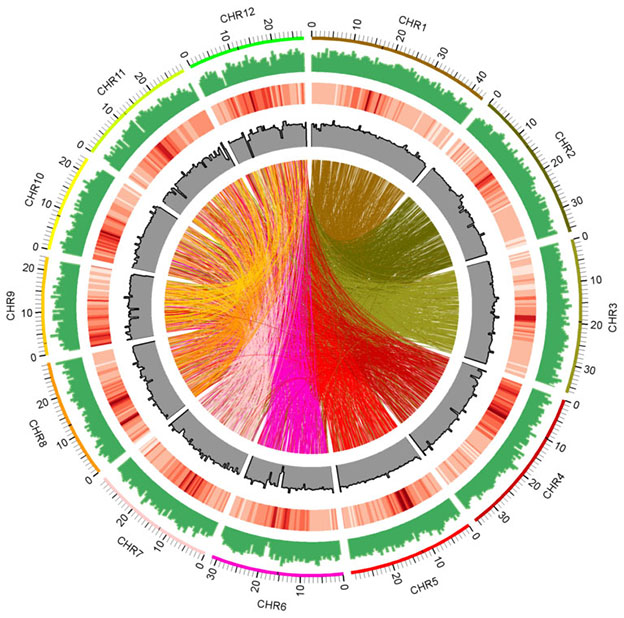
The Science
Researchers produced a database of mutations in an important grass crop: rice. To induce mutations they used fast-neutron irradiation. High-energy neutrons induce a wide variety of genetic mutations. Resequencing the 1,504 mutants allowed the researchers to identify structural variants and mutations. The results provide an invaluable resource to improve candidate bioenergy crops, such as switchgrass.
The Impact
Boosting yields of bioenergy feedstock crops such as grasses requires knowing in detail how enzymes and proteins synthesize plant cell walls to modify the processes and the composition. The mutant database offers a functional genomics resource for grass models. These models stand in for the biofuel crop switchgrass. The models are involved in biofuel studies to understand the challenges around plant cell wall synthesis. Until now, mutant collections for grass models have lagged behind those available for the Arabidopsis (flowering plants related to mustard and cabbage) model system.
Summary
For more than half of the world's population, rice is the primary staple crop. As a grass, it is a close relative of the candidate bioenergy feedstock switchgrass. The research team assembled the first major large-scale collection of mutations for grass models. Scientists at the University of California, Davis led the team, which includes researchers at the U.S. Department of Energy Joint Genome Institute (DOE JGI), a DOE Office of Science user facility, and the Joint Bioenergy Institute (JBEI), a DOE Bioenergy Research Center. They used the model rice cultivar Kitaake (Oryza sativa L. ssp. japonica), and compared the genes against the reference rice genome of another japonica subspecies called Nipponbare available on the DOE JGI Plant Portal Phytozome.
Through fast-neutron irradiation, the time-consuming procedures involving plant transformation or tissue culture were bypassed, allowing for faster development of rice mutant collections. The DOE JGI resequenced 1,504 rice mutants and identified structural variants and mutations. The work follows a pilot, genome-wide study begun two years ago, in which 41 rice mutants were sequenced and analyzed to identify mutations and structural variants. This new, large-scale collection of more than 90,000 mutations affecting nearly 60 percent of all rice genes is now available on a publicly accessible database called KitBase. The database is a comprehensive resource that will allow researchers to quickly identify rice lines with mutations in specific genes and to characterize gene function. Among other uses, the collection will allow bioenergy researchers to quickly identify mutations involved in cell wall biosynthesis, critical for increasing plant yields.
Contact
PM Contact
Daniel Drell, Ph.D.
Program Manager
Biological Systems Sciences Division
Office of Biological and Environmental Research
Office of Science
U.S. Department of Energy
daniel.drell@science.doe.gov
Kent Peters, Ph.D.
Program Manager
Biological Systems Sciences Division
Office of Biological and Environmental Research
Office of Science
U.S. Department of Energy
kent.peters@science.doe.gov
PI Contact
Jeremy Schmutz
Plant Program Head
DOE Joint Genome Institute
jschmutz@hudsonalpha.com
Pamela Ronald
University of California, Davis
pcronald@ucdavis.edu
Funding
Work was conducted by the U.S. Department of Energy (DOE) Joint Genome Institute, a DOE Office of Science user facility and the DOE Joint BioEnergy Institute (contract DE-AC02-05CH11231). This work was also supported by the National Institutes of Health and the National Science Foundation.
Publications
G. Li, R. Jain, M. Chern, N.T. Pham, J.A. Martin, T. Wei, W.S. Schackwitz, A.M. Lipzen, P.Q. Duong, K.C. Jones, L. Jiang, D. Ruan, D. Bauer, Y. Peng, K.W. Barry, J. Schmutz, and P.C. Ronald. "The sequences of 1,504 mutants in the model rice variety Kitaake facilitate rapid functional genomic studies. " The Plant Cell, (2017). [DOI: 10.1105/tpc.17.00443]
" The Plant Cell, (2017). [DOI: 10.1105/tpc.17.00443]
Related Links
- Lawrence Berkeley National Laboratory news release: A Whole-Genome Sequenced Rice Mutant Resource for the Study of Biofuel Feedstocks

- Plant Cell highlight: Technology Turbocharges Functional Genomics

- Kitaake Rice Mutant Database: KitBase

- Crop Genetics Innovation

- Joint Genome Institute's Plant Portal Phytozome: Quick search offers options to search for nipponbare/japonica subspecies of Oryza sativa
- Joint BioEnergy Institute: Feedstocks Division

- Joint Genome Institute science highlight: Grasses: The Secrets Behind Their Stomatal Success
Highlight Categories
Performer/Facility: University, DOE Laboratory, SC User Facilities, BER User Facilities, JGI
Additional: Collaborations, Non-DOE Interagency Collaboration

