Computers learn from a combination of experimental and evolutionary data to enhance the function of useful proteins.
Biological and Environmental Research
April 14, 2022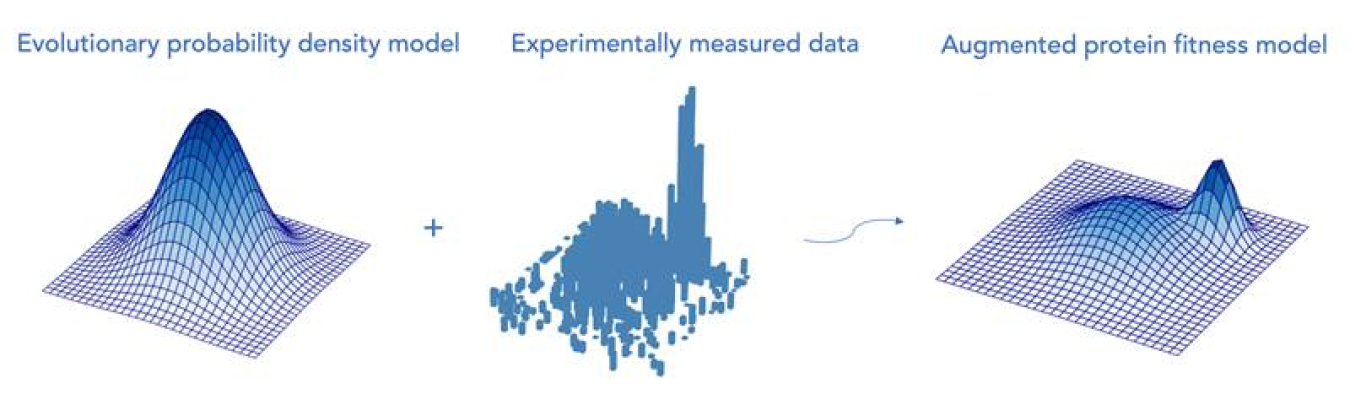
The Science
Proteins are integral components of all living organisms. They are composed of a sequence of building blocks called amino acids. That sequence determines their function, which can range from setting the structure of cells to regulating metabolism. Scientists can change a protein sequence and experimentally test if and how that change alters its function. However, there are too many possible amino acid sequence changes to test them all in the laboratory. Instead, researchers build highly complex computational models that predict protein function based on their amino acid sequence. This is critical for engineering proteins with novel functions. Scientists have now combined multiple machine learning approaches for building a simple predictive model that often works better than established, complex methods.
The Impact
Naturally occurring proteins serve many crucial functions in maintaining life. But scientists can also engineer natural proteins for desired purposes such as gene editing and the synthesis of valuable chemicals. This new combined modeling approach to predict protein function will aid in the design and engineering of novel proteins. This approach will allow scientists to easily redesign proteins for a huge range of applications such as new enzymes to convert plant matter into biofuels or bioproducts or to create new biomaterials.
Summary
Scientists have several approaches to predict functional properties of a given protein that use the protein’s amino acid sequence to build a computational model. Scientists create such models employing both classical statistical methods and modern-day machine learning computational approaches. One of those statistical methods, called regression analysis, associates a given amino acid sequence with an experimentally measured functional property of a protein. To increase the amount of data available to make functional predictions for a protein, researchers include sequences of evolutionarily-related proteins as additional input. In general, those evolutionarily-related proteins are likely to share the property of the protein of interest, albeit often without direct experimental evidence. Researchers use a machine learning modeling approach based on the statistical properties of those sequences. In the study highlighted here, researchers combined regression analysis and evolutionary data to propose a simple, effective machine learning approach. The researchers found that this simple combination approach is competitive with, and often outperforms, more sophisticated methods.
Contact
Chloe Hsu
University of California, Berkeley
chloehsu@berkeley.edu
Jennifer Listgarten
University of California, Berkeley
jennl@berkeley.edu
Funding
Partial support was provided by the Department of Energy Office of Science, Office of Biological and Environmental Research, Genomic Science Program, by Lawrence Livermore National Laboratory’s Secure Biosystems Design Scientific Focus Area, by the Chan Zuckerberg Investigator program, and by C3.ai. This material is also based upon work supported by the National Library of Medicine of the National Institutes of Health and the National Science Foundation Graduate Research Fellowship Program.
Publications
Hsu, C., Nisonoff, H., Fannjiang, C. and Listgarten, J., Learning Protein Fitness Models from Evolutionary and Assay-Labeled Data. Nature Biotechnology (2022).[DOI: 10.1038/s41587-021-01146-5]

