Bioprose Blog: LANL is developing genetic engineering tools that would allow algae to grow faster and accumulate more biomass for biofuel production.
Alternative Fuels and Feedstocks Office
January 10, 2019
Author: Amanda Barry, Algae Program Laboratory Relationship Manager, Los Alamos National Laboratory
Read Amanda's bio ►
Meet the other bloggers ►
Return to Bioprose blog ►
Bioprose Blog
Algae, like many things in life, should not be judged by their appearances. When we talk about algae, we’re really talking about a diverse group of organisms that thrive in a plethora of environments, from seawater to freshwater, from pulp mill ponds to hot springs. Some are strict phototrophs and can only use light for energy, whereas others can consume organic carbon, like raw plants, for growth. Algae are present in 4 of the 5 eukaryotic supergroups, making some of them as distinct from each other as we are from them. However, to the naked eye—and even under a microscope to a trained individual—algae look very similar to each other and it has only been through deep sequencing that the extent of their diversity has been revealed.
In the last ten years, available algae genomes (DNA) and the total number of transcriptomes (RNA) have increased significantly; many of these sequencing efforts were funded by the U.S. Department of Energy's Bioenergy Technologies Office (BETO). Still, there are many species of algae that have not been investigated (Figure 1), and, as Los Alamos National Laboratory (LANL) scientists recently discovered, there is still much to learn about the algae we thought we already understood.
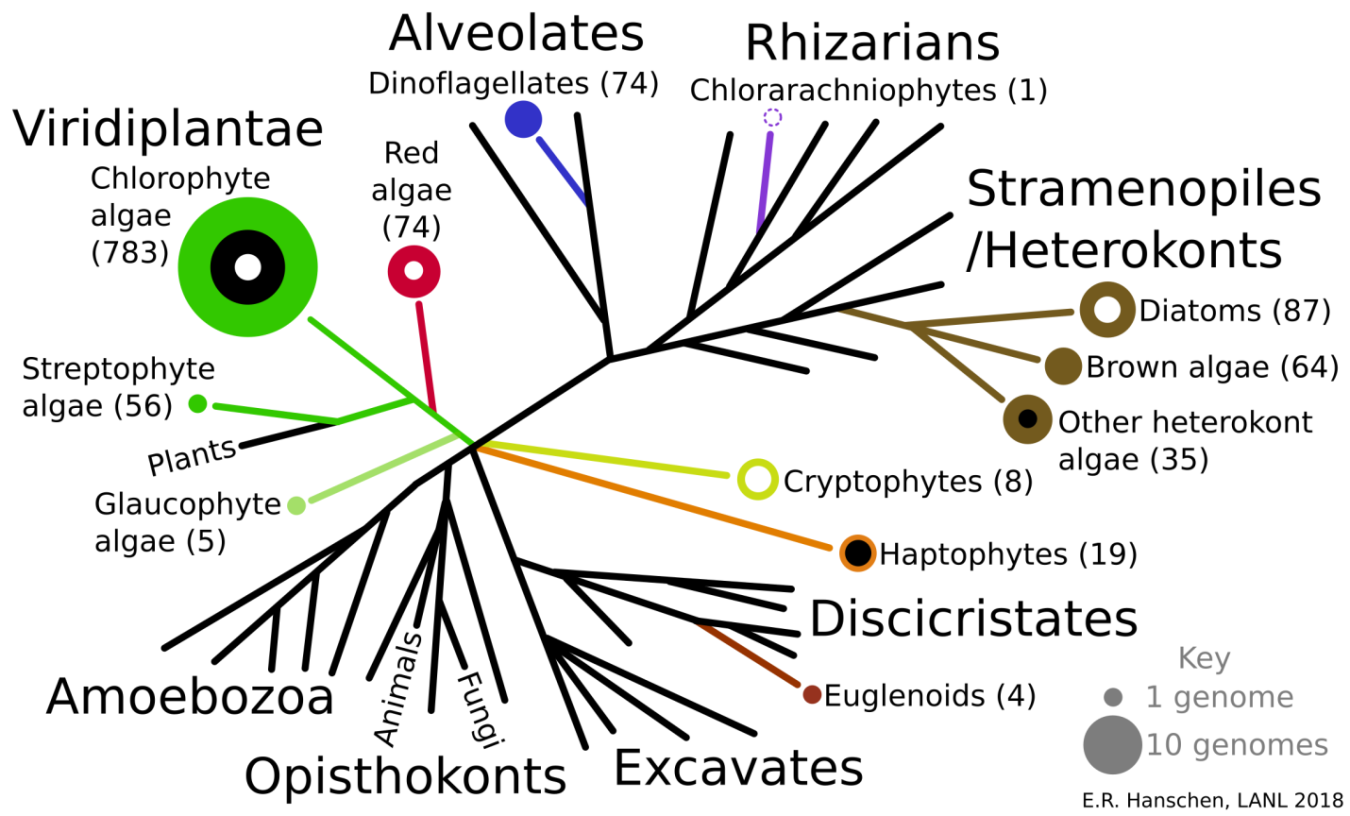
YOU CAN’T JUDGE A GREEN BALL BY ITS MORPHOLOGY
In a recent project funded by the Advanced Algal Systems (AAS) Program, researchers from LANL and the New Mexico Consortium are exploring the biofuel candidate Chlorella sorokiniana, developing genetic engineering tools in order to introduce novel traits that would allow the algae to grow faster and accumulate more biomass for biofuel production.
In order to improve growth, the team attempted to transform or change one of the Chlorella sorokiniana cultures by adding specific genes. However, they quickly found that some cultures grown from single colonies were amenable to genetic transformation, able to express the gene-of-interest, whereas others were not. This was a surprising clue that the C. sorokiniana culture they were studying might not be just one species.
In order to test this hypothesis, the LANL scientists characterized single algal colonies, discovering that some single colonies behaved differently than others. For example, some colonies exhibited the ability to degrade cellulose (an ability that could allow the algae to grow on alternative carbon substrates), while others had higher antibiotic tolerance (a trait important in genetic engineering) and salt resistance (a characteristic that may enable an algae to grow in brackish water). Under a microscope, they all appeared as little green balls and preliminary DNA analysis had indicated they were indeed a homogenous culture of Chlorella sorokiniana. When the LANL team fully sequenced the genomes of several single colonies, they found the culture was actually a mix of two strains of Chlorella sorokiniana, now named '1228 and 1230'. Although certain genetic sequences were identical, others varied significantly, resulting in two phenotypically and genetically distinct algae. The LANL team published a paper outlining the genetic characterization of these strains and a comparison of their genomes to a third Chlorella sorokiniana (1412) in “Genomic characterization reveals significant divergence within Chlorella sorokiniana (Chlorellales, Trebouxiophyceae)” by Hovde and Hanschen et al., in Algal Research (Figure 2).
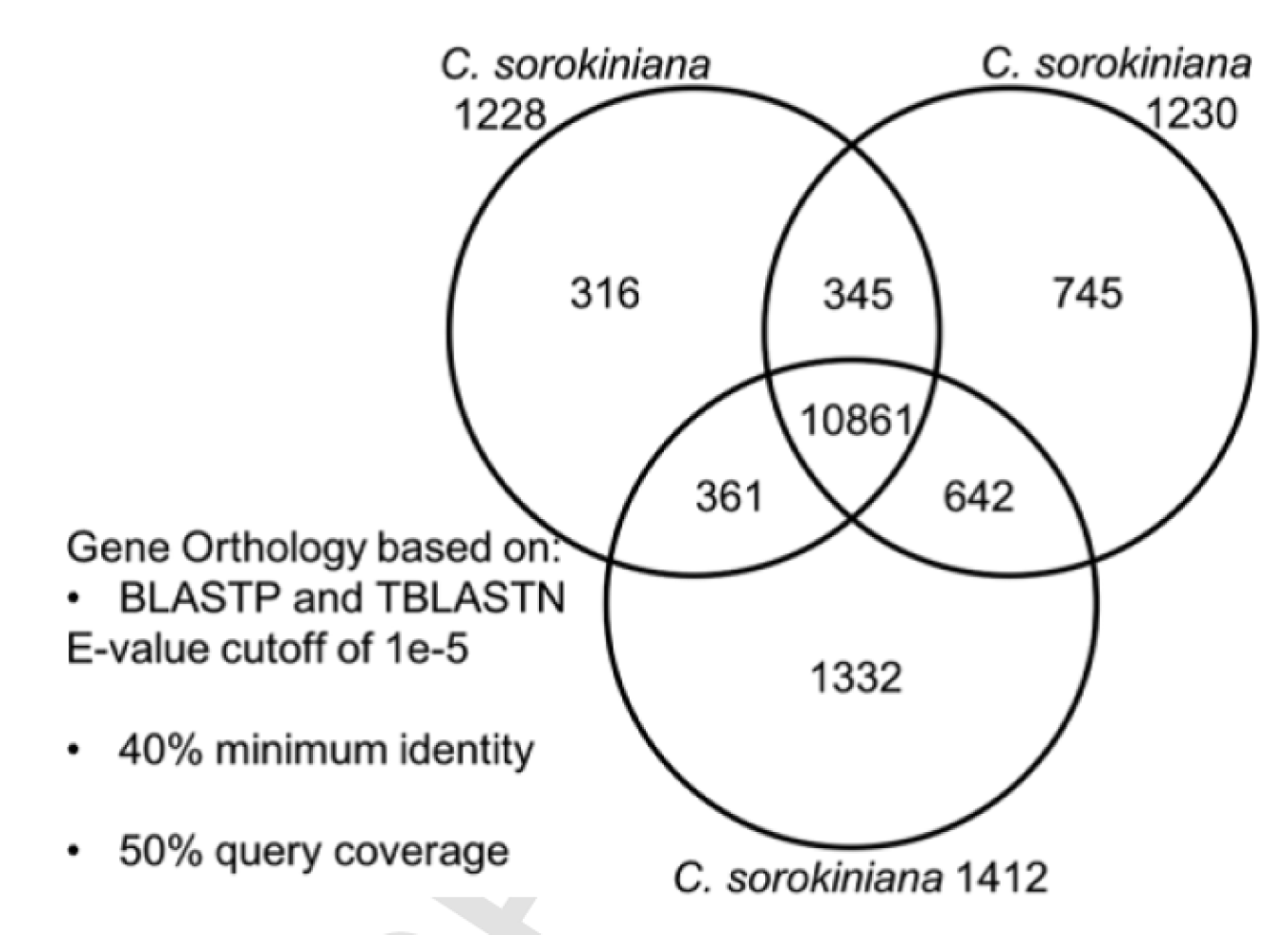
TAPPING INTO DIVERSITY
Tapping into this algal diversity to discover advantageous traits and marketable products is one of the ways researchers at LANL are working to make biofuels an economic reality. Other efforts at LANL to explore algal diversity for biofuels include exploring plant carbon utilization, identifying carbon and nitrogen utilization pathways in uncharacterized algal strains, examining the ecology of algal ponds, and participating in the Algae DISCVR (Development of Integrated Screening, Cultivar Optimization, and Validation Research) Project, a multi-lab collaboration that seeks to create a platform for the screening and characterization of algae strains under outdoor-relevant conditions for biofuel production. These efforts are facilitating BETO’s goals to achieve renewable and economical algae biofuels and bioproducts.
Amanda Barry

Dr. Amanda Barry is the R&D Manager for the Molecular and Microbiology Department at Sandia National Labs in Albuquerque. Previously, she worked at Los Alamos National Lab as a scientist and as the Algae Program Laboratory Relationship Manager. Dr. Barry's primary research focus is in algal biochemistry and the genetic manipulation of algae for economical biofuel production. From 2015-2017, Dr. Barry served on an Intergovernmental Personnel Act assignment at the U.S. Department of Energy, supporting the Advanced Algal Systems program in the Bioenergy Technologies Office (BETO) as Scientist/Technology Development. She co-authored the National Algal Biofuels Technology Review and served as Technical Lead for the verification of BETO-funded competitive awards. Dr. Barry received B.Sc. degrees in Biochemistry & Biophysics and in Bioresource Research from Oregon State University and a M.Sc. in Bacteriology from the University of Wisconsin-Madison. Her Ph.D. is from the Oregon Health and Science University in Biochemistry and Molecular Biology.
Meet our other bloggers ►
Return to Bioprose blog ►
MOST RECENT
-
 Bioprose Blog: LANL is developing genetic engineering tools that would allow algae to grow faster and accumulate more biomass for biofuel production.
Bioprose Blog: LANL is developing genetic engineering tools that would allow algae to grow faster and accumulate more biomass for biofuel production.


