High resolution isotope analysis of the algal microbiome identifies ecological strategies not predicted by genome content.
Biological and Environmental Research
February 20, 2024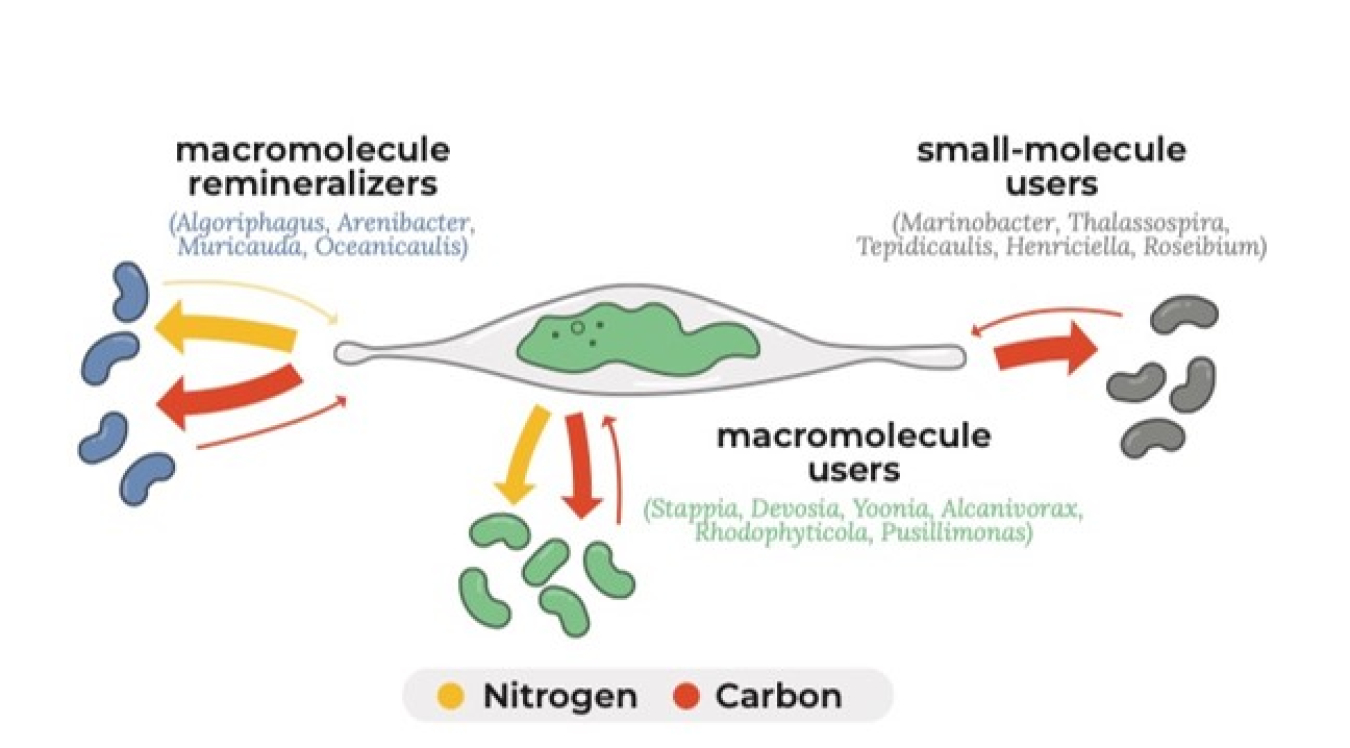
The Science
The microorganisms in a community—called a microbiome—interact in complex ways. In this project, researchers quantified the activity in the microbiome associated with tiny photosynthesizing algae. The team investigated how different members of this microbiome can process carbon and nitrogen from algal cells, then exchange some of that carbon and nitrogen back to the algae. They used isotopes and a high-resolution imaging mass spectrometer called NanoSIMS to quantify these exchanges at the level of single cells. This allowed them to identify different strategies that microbes can use in these microbiomes.
The Impact
Microscopic algae in water are responsible for roughly 50% of the photosynthesis that converts carbon dioxide from the Earth’s atmosphere into organic carbon. These organisms can be harnessed to grow as crops to produce carbon neutral fuels or even to bury carbon. Microbes such as bacteria are mainly responsible for breaking down this algal carbon and releasing it to the atmosphere. Understanding these microbes will help scientists predict and eventually alter the flux of carbon back to the atmosphere.
Summary
Bacterial remineralization of algal organic matter fuels algal growth, but scientists rarely quantify this remineralization. Consequently, researchers cannot predict whether some bacteria may provide more remineralized nutrients to algae than others. In this work, a team led by Lawrence Livermore National Laboratory with support from Pacific Northwest National Laboratory quantified bacterial incorporation of algal-derived complex dissolved organic carbon and nitrogen and algal incorporation of remineralized carbon and nitrogen. They examined this in 15 bacterial co-cultures growing with the diatom Phaeodactylum tricornutum at the single-cell level using isotope tracing and NanoSIMS. They found unexpected strain-to-strain and cell-to-cell variability in net carbon and nitrogen incorporation, including non-ubiquitous complex organic nitrogen utilization and remineralization.
The researchers then used these data to identify three distinct functional guilds of metabolic interactions. They termed these guilds macromolecule remineralizers, macromolecule users, and small-molecule users, the latter exhibiting efficient growth under low carbon availability. These functional guilds were not linked to evolutionary history and could not be elucidated strictly from metabolic capacity as predicted by comparative genomics. This highlights the need for direct activity-based measurements in ecological studies of microbial metabolic interactions.
Contact
Xavier Mayali
Lawrence Livermore National Laboratory
mayali1@llnl.gov
Funding
This work was funded by the Department of Energy Office of Science, Division of Biological and Environmental Research, Genomic Sciences Program under the microBiospheres Scientific Focus Area.
Publications
Mayali, X., et al., Single-cell isotope tracing reveals functional guilds of bacteria associated with the diatom Phaeodactylum tricornutum. Nature Communications 14, 5642 (2023). [DOI: 10.1038/s41467-023-41179-9]
Related Links
Not all bacteria are created equal, Lawrence Livermore National Laboratory News
The µBiospheres Scientific Focus Area research program at Lawrence Livermore National Laboratory

