Scientists at Los Alamos National Laboratory (LANL) have developed biosensing technology that can help address the problem of plastic waste, specifically nylon-based waste such as textiles, carpets, tires, and fishing nets. These and other products are...
Alternative Fuels and Feedstocks Office
October 25, 2023Dr. Sheila Van Cuyk

Dr. Sheila Van Cuyk is the Laboratory Relationship Manager for Bioenergy Technology Office programs at Los Alamos National Laboratory (LANL). She is a scientist in the Bioscience Division, LANL’s Biofuels Program Manager in the Applied Energy Program Office, and a National Security and Defense Program Manager for Global Security.
Sheila has a background in molecular biology and environmental engineering and has over 15 years of experience developing innovative interdisciplinary solutions to complex problems. Prior to her current role, she fulfilled an Intergovernmental Personnel Act billet as a Program Manager at the Department of Homeland Security, Science and Technology Directorate working in the areas of biological threat detection and biosurveillance.
Sheila completed her M.S. and Ph.D. in environmental engineering from the Colorado School of Mines and her B.S. in biology from the College of William and Mary.
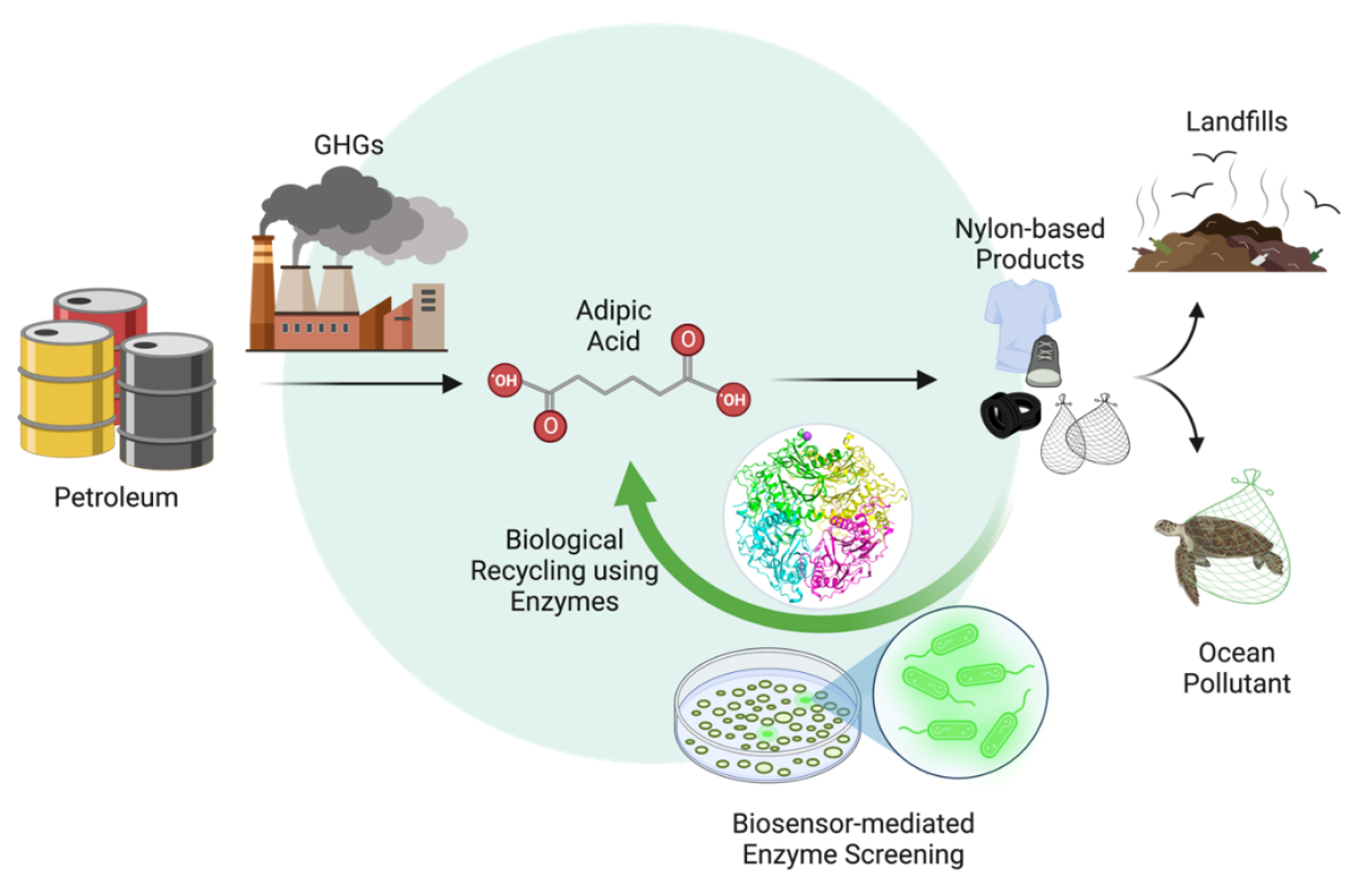
Scientists at Los Alamos National Laboratory (LANL) have developed biosensing technology that can help address the problem of plastic waste, specifically nylon-based waste such as textiles, carpets, tires, and fishing nets. These and other products are typically made from nylon-66, a common, very tough nylon made of a polymer building block called adipic acid (ADA). Unfortunately, the petroleum-based production of ADA releases nitrous oxide into the environment. Degrading, or depolymerizing, nylon using biocatalysts could pave the way to reducing nylon waste in landfills and oceans and enable recycling nylon building blocks into new products without the release of nitrous oxide.
The research, published in ACS Synthetic Biology, tackles an important challenge in this area: finding biocatalysts—typically enzymes or whole cell microbes— for anthropogenic molecules and optimizing them for industrial use. To find or engineer a biocatalyst that degrades nylon into its ADA building blocks, the scientists needed a way to detect the ADA molecules to confirm the biocatalyst had worked. The award-winning whole cell biosensor technology developed at LANL can rapidly screen candidate biocatalysts. The biosensor uses a bacterial cell host, such as Pseudomonas putida, and includes two target-specific recognition proteins: a transporter and a transcription factor.
“Our biosensor uses a sensor-reporter system to confirm when a candidate enzyme is successful,” explains LANL biologist Dr. Ramesh Jha, who participated in the U.S. Department of Energy (DOE) Bioenergy Technologies Office (BETO) research. “For example, we can test several nylon-degrading enzyme candidates, each in an individual biosensor cell. The enzyme seeps out of the cell and breaks down nylon polymers into ADA building block molecules, which are then transported back into the biosensor cells. The ADA that re-enters the cell binds to a transcription factor, which triggers production of green fluorescence protein, or GFP. The green glow of the GFP tells us the enzyme was active, and the intensity of the glow gives us information on the enzyme’s efficiency.”
As part of the Bio-Optimized Technologies to keep Thermoplastics out of Landfills and the Environment (BOTTLE) consortium, funded by BETO and DOE’s Advanced Materials & Manufacturing Technologies Office, and the BETO-funded Agile BioFoundry consortium, the Los Alamos scientists wanted to use their biosensor to degrade nylon-66. However, for anthropogenic molecules such as nylon and its building block molecule, ADA, Jha and his team knew it would be difficult to find naturally occurring proteins to recognize these targets. It was essentially a “Catch-22”-type situation where engineering and validating the two target recognition proteins, transporter and transcription factor, were dependent on each other: establishing a transporter needed a sensitive intracellular detector such as a biosensor and optimizing a whole cell biosensor depended on an active transporter.
To address this frustrating circular problem, the LANL team looked to a previous biosensor they had made for the naturally occurring molecule, muconate, which is chemically similar to ADA. With an available transporter for muconate in nature, the team hypothesized that a muconate transporter might be able to transport ADA too—and it did! However, the key obstacle was that the transporter was “toxic” to the Pseudomonas putida bacterial host when produced at a certain level. Using their existing muconate biosensor, the team tuned the transporter expression to below the toxicity level so that the transporter would remain functional for many P. putida generations.
The next step was to engineer a highly specific transcription factor for ADA. The team used their computational modeling expertise and a focused, mutational library to design 20 million variants of a transcription factor, called PcaR, that could theoretically bind to ADA to initiate production of GFP reporter. The scientists coupled the biosensor to an instrument called a flow cytometer that detects the green fluorescence and sorts the cells based on the fluorescence intensity—this enabled high-throughput screening of all 20 million variants. After five rounds of fluorescence-activated cell screening, three variants of PcaR were isolated that responded in various degrees to ADA. “It took us four weeks to build a biosensor for this molecule. The approach that combined the computationally informed library design and high throughput screening using flow cytometry greatly accelerated the biosensor development process and made it possible to evaluate millions of variants in a timely manner,” says Dr. Sang-Min Shin, a member of the Los Alamos team.
The Los Alamos whole cell biosensor is the first-ever ADA biosensor in P. putida. “We are actively using the biosensor to discover and engineer efficient nylon-degrading enzymes for “green” recycling of the building blocks or upcycling into bioadvantaged products” says LANL biochemist Dr. Taraka Dale, who led the R&D team for biosensor development.
Dr. Sheila Van Cuyk is the Laboratory Relationship Manager for Bioenergy Technology Office programs at Los Alamos National Laboratory (LANL).


