Scientists discover a mechanism for plant-microbe interactions.
Biological and Environmental Research
October 18, 2023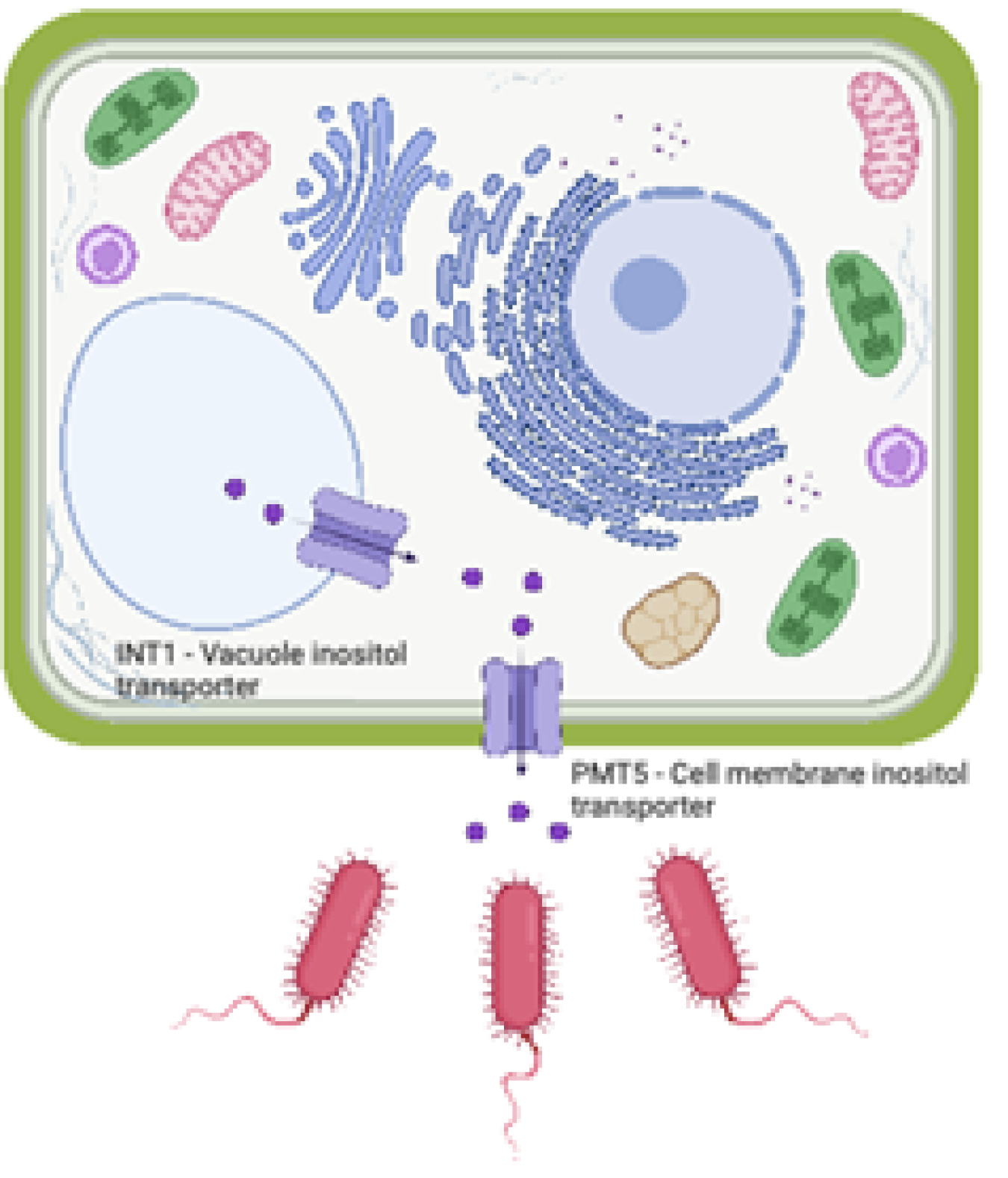
The Science
Microorganisms that live on or in plant tissues form what is known as a plant microbiome. This interface plays an important role in plants’ survival due to the existence of beneficial microorganisms. Plants grown in distinct environments can have similar microbiomes or can change over time depending on environmental factors. This complex microbial community assembles and changes by exchanging signals between the host and microbes. In this study, researchers gathered and filtered a large amount of data using a combination of computational approaches to identify new mechanisms. They then used experiments to validate these mechanisms. This data mining led to the discovery of a host transport mechanism and a chemical signal that influences beneficial bacterial colonization of plants’ roots.
The Impact
Across the tree of life, microbiomes come together through a complex dialogue between host plants and their microbial partners. This research adds to scientific knowledge about these dialogues. The study highlighted a unique role for how host plants transport a molecule that influences how microbes colonize microbiomes. The study used an experimental approach to filter large datasets. This approach will improve researchers’ ability to identify new chemical signals. Understanding signaling molecules that are involved with increasing colonization by beneficial microbes will help scientists examine new ways to help plants resist pathogens and reduce the effects of environmental stress.
Summary
To be able to identify microbial taxa in any sequence dataset, researchers built kmer profiles from every publicly available, sequenced genome. Using these kmer profiles with their ParaKraken codebase on the Summit supercomputer at the Oak Ridge National Laboratory, they analyzed meta-transcriptomic sequencing data from leaf and xylem tissue from approximately 500 Populus trichocarpa genotypes grown in a common garden. This approach allowed the researchers to detect thousands of species of microbes living in these plant tissues. They used the abundance of each species as a phenotype for a genome-wide association study to determine which plant genes were likely affecting the colonization of each microbial species. This resulted in a rich view of the processes that the host plant uses to select for specific microbial species in its microbiome. The researchers found that two different microbial species were both affected by both of the myo-inositol transporters in plant xylem (stem) tissue. To study this finding further, they used the model plant species Arabidopsis thaliana (a small mustard weed) to do so. They used existing lines of Arabidopsis in which these myo-inositol transporters had been deleted. The researchers measured the colonization levels of Arabidopsis seedling roots in the knockout lines versus a control that contained the genes in a laboratory-based assay where the seedlings were grown on agar plates.
They found that the Arabidopsis lines without myo-inositol transporters had significantly reduced levels of colonization. Furthermore, colonization levels were restored when the researchers added myo-inositol to the growth medium in the agar plates. They found that the bacterial colonization is controlled by the same genes in both the stem tissue in trees grown under field conditions and in the roots of mustard weed seedlings grown in the laboratory. This remarkable finding and points to the strong conservation of this mechanism across very different types of plants. The researchers further investigated the mode of action of myo-inositol, which is known to be an internal signaling molecule in plants. Surprisingly, they found that knocking out genes in the plant signaling cascade did not affect colonization levels in Arabidopsis roots. Myo-inositol is a type of sugar that some bacteria can use as a food source, so the researchers knocked out the catabolic pathway for myo-inositol in the bacteria and found that this also did not affect colonization. However, the researchers did discover that myo-inositol significantly affected the motility (swimming capability) of the bacteria. Thus, it appears that plants are using myo-inositol in a role that has never been studied before, specifically as a cross-kingdom signaling molecule. The plant thus appears to be pumping myo-inositol out of its tissues to trigger specific bacteria to swim towards plant roots and colonize them. In summary, researchers used species abundances in the Populus microbiome as phenotypes for genome-wide association studies (GWAS) to identify plant genes controlling colonization. They subsequently used experimental gene knockout assays in seedlings of the model plant Arabidopsis thaliana for host gene validation. The research discovered and confirmed a conserved role for the transport of the plant metabolite myo-inositol as a eukaryotic-derived signaling molecule to modulate microbial activities.
Contact
Dan Jacobson
jacobsonda@ornl.gov
Oak Ridge National Laboratory
Funding
This work was sponsored by the Department of Energy Office of Science, Biological and Environmental Research program’s Genomic Science Program as part of the Plant-Microbe Interfaces Science Focus Area and the Center for Bioenergy Innovation. The Poplar GWAS Project used resources of the Oak Ridge Leadership Computing Facility and the Compute and Data Environment for Science at Oak Ridge National Laboratory. This work was also supported by the Science Alliance Joint Directed Research and Development Funding at Oak Ridge National Laboratory, the Tennessee Plant Research Center, and the National Science Foundation.
Publications
O'Banion, B.S., et al., Plant myo-inositol transport influences bacterial colonization phenotypes. Current Biology 33, 1-14 (2023). [DOI: 10.1016/j.cub.2023.06.057]

